Delta Variant? There Are Others. Milford Molecular Diagnostics Laboratory Routinely Diagnoses Variants of SARS-CoV-2, Including Alpha, Beta, Gamma, Delta, Epsilon, Eta, Iota, Kappa and Lambda Variants
Delta Variant? There Are Others. Milford Molecular Diagnostics Laboratory Routinely Diagnoses Variants of SARS-CoV-2, Including Alpha, Beta, Gamma, Delta, Epsilon, Eta, Iota, Kappa and Lambda Variants
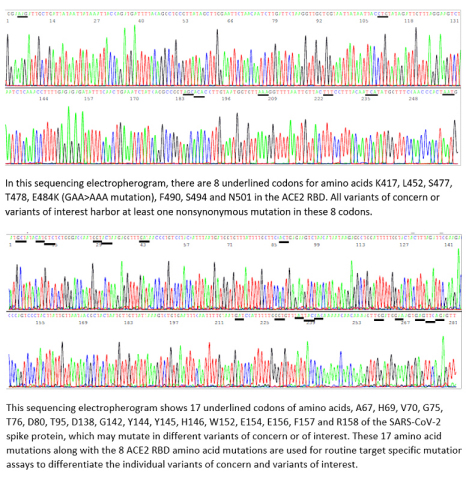
Target Specific Mutation Assays Are Based on Analysis of Two Unique Segments of S Protein gene
MILFORD, Conn.--(BUSINESS WIRE)--Currently circulating in the United States, the SARS-CoV-2 variants of concern are the Alpha, Beta, Gamma and Delta, and the variants of interest are the Epsilon, Eta, Iota, Kappa and Lambda. The Delta variant makes up 83% of new COVID-19 cases, said Dr. Sin Hang Lee, director of Milford Molecular Diagnostics, summarizing recently published sequence-based surveillance data.
“However,” he added, “there exists no known diagnostic method for these variants in the U.S. for individual patient samples outside of Milford Diagnostics Laboratory. This is because these variants, especially the Delta variant, carry distinctive biological markers that the commercially available COVID tests from academic institutes and medical giants are unable to detect. It requires target specific mutation assays to diagnose these variants in all positive samples.”
Dr. Lee explained the target specific mutation assays developed in his CLIA-certified laboratory as follows:
According to the publications from the CDC and the WHO, the distinctive biological markers for these variants are based on their unique sets of spike protein substitutions. A concise set is summarized below.
Alpha 69del, 70del, 144del, N501Y (B.1.1.7 U.K.)
Beta D80A, K417N, E484K, N501Y (B.1.351 South Africa)
Gamma D138Y, K417T, E484K, N501Y (P.1 Japan/Brazil)
Delta T95I, G142D, E156-, F157-, R158G, L452R, T478K (B.1.617.2 India)
Delta plus T95I, G142D, E156-, F157-, R158G, K417N, L452R, T478K (B.1.617.2.1 India)
Epsilon L452R (B.1.427 California)
Epsilon W152C, L452R (B.1.429 California)
Eta A67V, 69del, 70del, 144del, E484K (B.1.525 U. K./Nigeria)
Iota T95I, E484K (B.1.526 New York)
Kappa G142D, E154K, L452R, E484Q (B.1.617.1 India)
Kappa G142D, L452R, E484Q (B.1.617.3 India)
Lambda G75V, T76I, L452Q, F490S (C.37 Peru)
The variants, with their respective WHO labels in boldfaced Greek letter names, are on the left, followed by their distinctive specific amino acid mutations, Pango lineage and locations of first identification. The 8 invariably mutated amino acids of the angiotensin-converting enzyme 2 (ACE2) receptor binding domain (RBD) of the spike (S) protein are underlined. All variants of concern or of interest must have at least one of these 8 mutated amino acids. Mutations of the S protein amino acids without an underline are used to determine the individual variant. For example, to diagnose a Delta variant the laboratory must show at least 7 specific amino acid mutations, including two amino acid deletions and 5 nonsynonymous single nucleotide mutations, causing 5 amino acid changes. To diagnose an Epsilon variant, the laboratory must show a solitary L452R amino acid mutation in the ACE2 RBD with or without a concomitant W152C mutation.
Milford Molecular Diagnostics Laboratory is CLIA-certified to perform nested RT-PCR/DNA sequencing for diagnosis of SARS-Co-V-2 and reflex spike protein gene sequencing for variant determination.
Individuals interested in target specific mutation assays for SARS-CoV-2 variant determination may visit the website at http://dnalymetest.com or contact Kevin Moore at the number below for more information.
Contacts
Media :
For Milford Molecular Diagnostics Laboratory, Milford, Conn.
Kevin Moore, 203-788-8497